« Overview

DNA. More than a genetic blueprint.
Using just four bases (A,C,G and T), DNA — a molecule ubiquitous
to life as we know it — acts as a storage device by encoding
genes and other blueprint sequences that can be inherited by future
generations. However, DNA is not limited to the role of information
carrier. Its very mention elicits thoughts of its famous double
helical structure. This structure is formed by two sequences that
hybridize together by forming bonds between complementary bases. The
A base will bond with a T base and, similarly, C will bond with G.
Using this knowledge, researchers can design sets of sequences that
will hybridize and self-assemble into shapes much more complex than
the double helix. This is the foundation of DNA nanotechnology.
Beyond silicon. Computing with Soup.
In addition to self-assembly into static structures, DNA
hybridization can be leveraged for creating dynamic systems that
change over time. This has naturally led to the exploration of
using DNA to perform computation. While the physical realization of
today's computer programs is in silicon hardware, scientists are
envisioning radically new ways of programming with bio-molecules
such as DNA that run in an energy-efficient manner at staggeringly
small scales. The advantage? A natural interface with biological
systems that can be implemented within a living cell. Imagine a
truly smart drug delivery system that only treats unhealthy cells.
Such a system must reason and can be implemented with a so-called
DNA Strand Displacement (DSD) system. In a DSD, DNA strands are
conceptually composed of different domains —a
particular sequence of bases. By a careful design of domains that
can hybridize to other domains, entire reaction cascades can be
implemented to perform complex logical computation within a
solution. This has been aptly described as computing with
soup.
Wetware Verification. Probabilistic Model Checking.
Correctly implementing traditional software or hardware is already a
difficult and error-prone task. Consider the additional challenges
when implementing a DNA based computer composed of millions or
billions of strands. At once, the strands form the hardware of the
system and their potential interactions comprise the software. In
any particular configuration of strands, or state of the system, any
potential binding of complementary domains leads to a new state.
Programmers only have control over the design of the strands. Can
they be certain strands interact in only desirable ways? As an end
goal of these systems is to run within living cells, safety and
correctness are of paramount concern. Yet, wet lab experimentation
is both time consuming and expensive and the scale of these systems
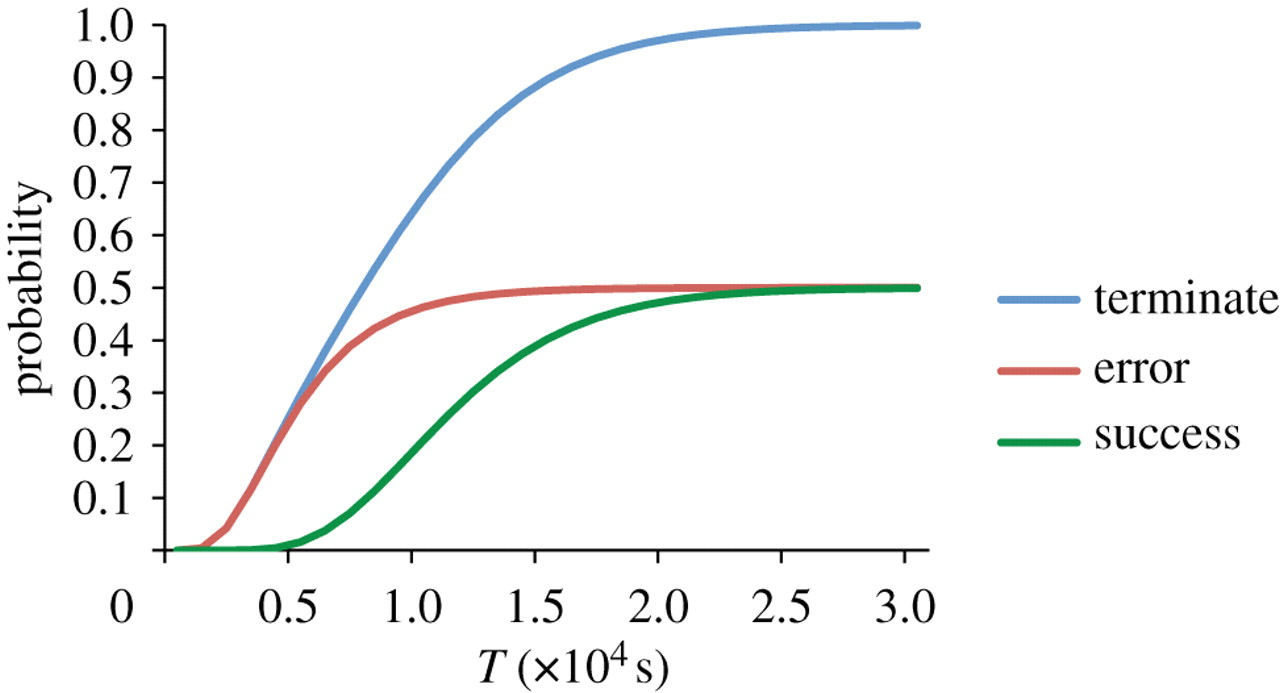
prohibits direct observation. Fortunately, quantitative
verification of these highly concurrent, probabilistic systems is
possible. The PRISM
model checking tool has already been used to great effect in
identifying bugs in these systems, and verifying their
expected behaviour. Work continues in this project to improve the
state-of-the-art for model checking these critical systems and the
unique challenges they present.
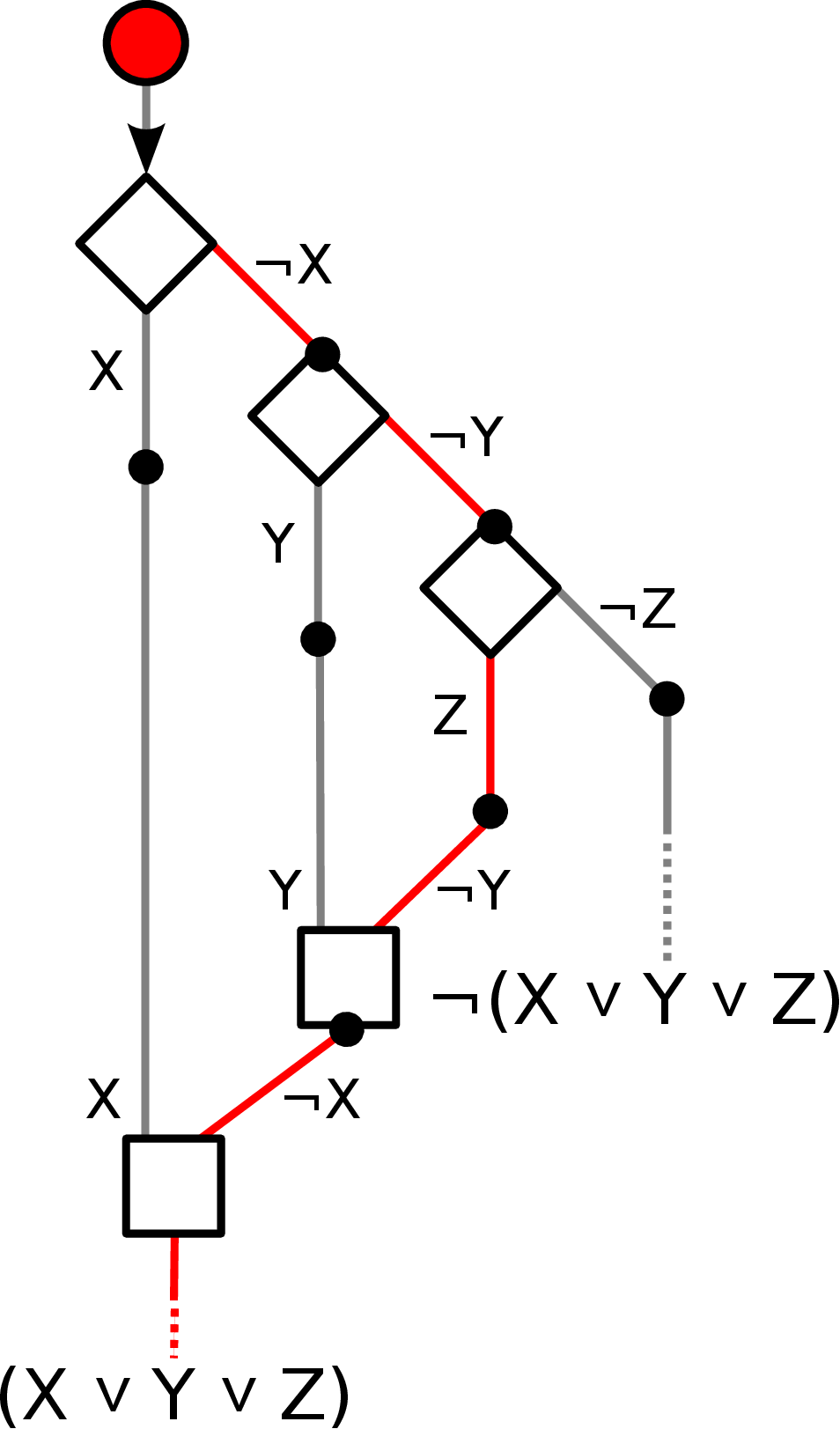
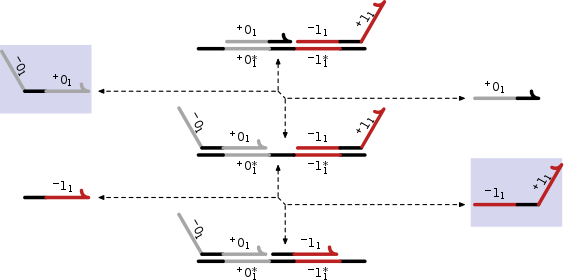
Walker systems. Expressing logic gates.
Various walkers systems have been constructed through clever manipulation of DNA.
These advanced motors can operate along a track made of DNA and can take directions at junctions.
Their function mimics the natural proteins Kinesin and Dynein, which transport cargo along
cytoskeletal microtubules inside the cell. Advances made in engineering these artificial systems
clear the path towards mechanical designs with more complicated functions, such as logic gates
or persistent memory made from DNA. Such DNA computers share natural interfaces with biological
inputs and could one day function as advanced bio-sensors. As the complexity increases, so does
the necessity of automated design tools for DNA systems. Model checking can analyse the reliability,
detect bugs and predict the performance of future experiments.
Sort by: date, type, title
18 publications:
-
[CKL16b]
L. Cardelli, M. Kwiatkowska and L. Laurenti.
A Stochastic Hybrid Approximation for Chemical Kinetics Based on the Linear Noise Approximation.
In Proc. 14th International Conference on Computational Methods in Systems Biology ({CMSB} 2016), Springer. To appear.
2016.
[pdf]
[bib]
-
[CKW16]
L. Cardelli, M. Kwiatkowska, M. Whitby.
Chemical Reaction Network Designs for Asynchronous Logic Circuits.
In Y. Rondelez and D. Woods (editors), Proc. 22nd International Conference on DNA Computing and Molecular Programming (DNA22), Springer. To appear.
2016.
[pdf]
[bib]
-
[CDP+16]
M. Ceska, F. Dannenberg, N. Paoletti, M. Kwiatkowska and L. Brim.
Precise Parameter Synthesis for Stochastic Biochemical Systems.
Acta Informatica, Springer. To appear.
2016.
[pdf]
[bib]
-
[Dan16]
F. Dannenberg.
Modelling and verification for DNA nanotechnology.
Ph.D. thesis, Department of Computer Science, University of Oxford.
2016.
[pdf]
[bib]
-
[BCKL16]
L. Bortolussi, L. Cardelli, M. Kwiatkowska and L. Laurenti.
Approximation of Probabilistic Reachability for Chemical Reaction Networks using the Linear Noise Approximation.
In Proc. 13th International Conference on Quantitative Evaluation of SysTems (QEST 2016), Springer. To appear.
2016.
[pdf]
[bib]
-
[CKL16a]
L. Cardelli, M. Kwiatkowska, L. Laurenti.
Programming Discrete Distributions with Chemical Reaction Networks.
In Y. Rondelez and D. Woods (editors), Proc. 22nd International Conference on DNA Computing and Molecular Programming (DNA22), Springer. To appear.
2016.
[pdf]
[bib]
-
[DDB+15]
F. Dannenberg, K. Dunn, J. Bath, M. Kwiatkowska, A. Turberfield, and T. Ouldridge.
Modelling DNA Origami Self-Assembly at the Domain Level.
Journal of Chemical Physics, 143(16).
2015.
[bib]
http://arxiv.org/abs/1509.03066
-
[CKL15a]
L. Cardelli, M. Kwiatkowska, L. Laurenti.
Stochastic Analysis of Chemical Reaction Networks Using Linear Noise Approximation.
In Proceedings of 13th Annual Conference on Computational Methods in Systems Biology (CMSB 2015), volume 9308 of LNCS, pages 3-7.
2015.
[pdf]
[bib]
-
[DKTT15]
F. Dannenberg, M. Kwiatkowska, C. Thachuk and A. J.Turberfield.
DNA walker circuits: Computational potential, design, and verification.
Natural Computing, 14(2), pages 195-211, Springer.
2015.
[pdf]
[bib]
-
[BK15]
B. Barbot, M. Kwiatkowska.
On Quantitative Modelling and Verification of DNA Walker Circuits Using Stochastic Petri Nets.
In Devillers, Raymond and Valmari, Antti (editors), Proc. 36th International Conference on Application and Theory of Petri Nets and Concurrency, volume 9115 of Lecture Notes in Computer Science, pages 1-32, Springer International Publishing.
2015.
[pdf]
[bib]
-
[DDO+15]
K. Dunn, F. Dannenberg, T. Ouldridge, M, Kwiatkowska, A. Turberfield, J. Bath.
Guiding the folding pathway of DNA origami.
Nature, 525, pages 82–86, Nature Publishing Group.
September 2015.
[bib]
http://dx.doi.org/10.1038/nature14860
-
[WHI15]
Whitby, M..
The Construction and Verification of Asynchronous Components Built from Chemical Reaction Networks.
Masters thesis, University of Oxford.
June 2015.
[pdf]
[bib]
-
[DHK15]
F. Dannenberg, E. M. Hahn and M. Kwiatkowska.
Computing Cumulative Rewards using Fast Adaptive Uniformisation.
In ACM Transactions on Modeling and Computer Simulation, Special Issue in Computational Methods in Systems Biology, Association for Computing Machinery (ACM). Article 9.
April 2015.
[pdf]
-
[DHK13]
F. Dannenberg, E. M. Hahn and M. Kwiatkowska.
Computing Cumulative Rewards using Fast Adaptive Uniformisation.
In Proc. 11th Conference on Computational Methods in Systems Biology (CMSB'13), volume 8130 of LNCS, pages 33-49, Springer.
2013.
[pdf]
[bib]
-
[DKTT13]
F. Dannenberg, M. Kwiatkowska, C. Thachuk and A. J. Turberfield.
DNA Walker Circuits: Computational Potential, Design, and Verification.
In Proc. 19th International Conference on DNA Computing and Molecular Programming (DNA 19), volume 8141 of LNCS, pages 31-45, Springer.
2013.
[pdf]
[bib]
Sort by: date, type, title
« Overview






